Abstract
Abstract: Life science organizations are increasingly using hackathons to bring communities together to tackle shared problems, teach skills, and develop new resources. In this study, we explored the potential benefits of hackathons for the biotechnology workforce education community by organizing two hackathons centered around developing research projects in antibody engineering—a practice widely employed in the biotechnology industry but uncommon in biotechnology education. To integrate antibody engineering into courses, instructors need protocols for both computational and laboratory methods. Developing and testing these protocols provides rich opportunities for undergraduate research, allowing students to learn industry-relevant skills and contribute to creating materials for the community. During the hackathons, teams of faculty, students, and industry partners collaborated to generate several new research projects. Each hackathon was only a few days, yet student participants reported benefits similar to those attributed to traditional undergraduate research experiences. We share lessons learned from these hackathons and provide insights for the workforce education community for hosting similar events.
Keywords: antibody, CURE, hackathon, SARS-CoV-2, machine learning, epitope, iCn3D, undergraduate research
© 2023 under the terms of the J ATE Open Access Publishing Agreement
Introduction
Undergraduate research enhances student retention and graduation rates, especially for minority students in STEM fields [1-3], but community college students often lack opportunities for such valuable experiences due to limited resources. With 38% of undergraduate students attending community college [4], it is important to find ways to address this disparity.
Course-based undergraduate research projects (CUREs) have been proposed as a method for increasing undergraduate research experiences (UREs). Including research projects in courses enables students to use scientific practices, make new discoveries, and contribute to knowledge [5, 6]. Implementing CUREs, however, requires funding, professional development for instructors, and course modifications.
In the context of college biotechnology and biomanufacturing programs, CUREs also need to align with learning goals designed to meet the demands of local industries. For simplicity, we refer to these programs as biotechnology programs in the rest of this article. Students in biotechnology programs require UREs or CUREs that offer opportunities to develop skills essential for the workforce. Research projects focused on antibody engineering can effectively address this need, as they involve both bioinformatics and laboratory techniques.
To develop antibody-engineering research projects while providing the professional development needed for implementation, we looked for a cost-effective platform that could be scalable, national, and informed by current industry needs and practices. We also wanted to start building a community of instructors with a shared knowledge base, both to improve sustainability and provide a way for instructors to get help.
Using Hackathons to develop CUREs for Antibody Engineering
We are piloting hackathons as a new approach to enlist the community in developing biotechnology-related undergraduate research experiences (UREs) and course-based undergraduate research projects (CUREs) that align with industry needs. Research projects in engineering antibodies are particularly relevant to the industry since they provide opportunities to engage in industry practices.
Hackathons are intensive, short-term events where teams collaborate to generate solutions or create prototypes. Hackathons have become increasingly popular in bioscience communities for fostering collaboration, inspiring creative thinking, building community, and tackling significant challenges. They are a regular feature at scientific conferences like BioIT World [7] and ISMB [8]. Government institutions, including the National Institutes of Health (NIH), routinely use hackathons to develop web applications and facilitate large-scale collaborations [9]. Since 2016, the NIH has organized over 40 hackathons with over 3000 participants, resulting in over 20 publications (Allissa Dillman, personal communication).
Despite their effectiveness, hackathons have not been explored in the ATE (Advanced Technological Education) community. We wanted to determine if hackathons could help us meet our goals of building community, providing professional development, and catalyzing the development of innovative projects around antibody engineering. Since collaboration is already the norm in biotech companies and a growing practice in the bioscience community [10], we also sought to foster teamwork and model a collaborative and supportive environment.
Why focus on Antibody Engineering?
Antibodies are one of the most important types of molecules that biotechnology programs address. These proteins, made by animal immune cells, bind to specific three-dimensional shapes (epitopes) on proteins and other molecules. By fusing an antibody-producing cell with a tumor cell, an immortal cell line capable of secreting identical antibodies (monoclonal antibodies) can be generated. Antibodies can also be produced by introducing plasmids with antibody genes into bacteria, yeast, and mammalian cells.
In biotechnology and biology research, antibodies are ubiquitous. In research, antibodies are used as reagents for detecting and/or purifying other molecules. In biotechnology, antibodies are made into drugs and diagnostic products such as home pregnancy tests [11] and tests for COVID-19 [12].
When it comes to therapeutics, monoclonal antibodies are the largest class of biopharmaceuticals on the market. Over 165 antibody-based drugs have either been approved by the FDA or EU regulatory agencies or were undergoing regulatory review in March 2023 [13]. All therapeutic antibodies have been engineered [14]. Daclizumab, one of the first engineered antibodies, received FDA approval in 1997 [15, 16]. This antibody was derived from mice but modified to resemble a human antibody by substituting mouse amino acids with their human counterparts, a process known as humanization. Humanization is used to minimize the risk of an immune response against the drug.
Antibody engineering encompasses making amino acid substitutions to improve physical characteristics like solubility, alterations in the binding site to improve specificity and affinity, conjugation with toxic molecules to enhance drug activity, and changes designed to optimize antibody manufacturing [14]. Additionally, many new antibody drugs are engineered to be bispecific, enabling them to bind two different antigens [17]. Engineered antibody genes can also be introduced into T cells as chimeric antigen receptors (CARs), designed to trigger an immune response against tumors.
Tools for Antibody Engineering
Web-based computational tools and databases have become valuable resources for structural biology research and education. Tools like iCn3D [18], the Immune Epitope Database (IEDB) [19], and SAbPred [20] offer molecular modeling, epitope analysis, and antibody structure prediction capabilities, respectively. These tools are freely accessible online and provide researchers with visualization and manipulation capabilities, as well as analysis and prediction tools. iCn3D, from the National Center for Biotechnology Information (NCBI, https://www.ncbi.nlm.nih.gov/), is a web-based molecular modeling tool for visualizing and manipulating molecular structures. IEDB offers analysis tools for studying the molecules recognized by antibodies [19]. SAbPred, developed by the Oxford Protein Informatics Group (OPIG), utilizes deep learning to predict antibody and nanobody structures, facilitating model generation, prediction, mutation analysis, and structure comparisons [20].
On the laboratory side, non-profit organizations like AddGene (Addgene.org) and government entities like the National Institute for Standards and Technology (NIST, NIST.gov) offer biological materials such as plasmids with antibody genes [21] and standardized cell lines [22] that can provide starting materials for antibody projects. AddGene has over 1362 plasmids with antibody genes, including genes for nanobodies. Nanobodies are stable, low molecular weight proteins (~15 kilodaltons), with a single protein chain derived from antibodies found in camels, llamas, and alpacas [23]. NIST recently developed a standardized version of the Chinese Hamster Ovary (CHO) cell line. CHO cells are the most common industrial system for manufacturing antibodies [24].
Not only are biological materials and computational tools available for educational use, but the process of antibody engineering offers multiple steps that can serve as starting points for research projects. Computational projects can involve molecular modeling, prediction, and plasmid design, while laboratory projects can explore purification methods, detection techniques, screening, assay development, specificity, binding strength, glycosylation, and more.
Computational projects also offer flexibility, as they can be carried out remotely, making them suitable for online courses or students unable to attend in-person classes. Furthermore, the low start-up costs for computational projects enable colleges with limited resources to engage students in cutting-edge research. Projects can also be divided into phases, with some work conducted in a physical lab and computational work carried out remotely.
Antibodies in Biotechnology Education
At least 364 US employers in 579 locations list antibodies as a key business area (https://biotech-careers.org/company-core-activity/antibodies), so it is not surprising that many biotech programs teach antibody-related laboratory skills. These include culturing mammalian cells, upstream processing (producing antibodies from CHO cells and monitoring fermentation), and downstream processing (antibody purification). These skills are important for biomanufacturing technicians who manufacture antibodies for therapeutics, diagnostics, and reagents. Analytical skills such as protein gel electrophoresis, protein assays, Western blots, enzyme-linked immunoassays (ELISAs), fluorescent antibody staining, and, more recently, flow cytometry [25] are also taught. These types of analytical skills are used by laboratory technicians in both biotechnology companies and research labs.
Antibody engineering, however, has not been part of two-year college biotech programs. Given the large number of companies that manufacture antibody-related products and the increasing use of antibody engineering in industry, it is important for instructors to learn more about these technologies. The rapid development of artificial intelligence and machine learning tools are revolutionizing the antibody development process. Soon it will be possible to create novel antibodies without immunizing animals.
If we are to use these powerful, new computational tools for antibody design in the classroom, instructors will need professional development. Others will need protocols and guidance for implementing laboratory techniques. Creating research projects from new materials and tools requires another level of learning, familiarity, and practice.
Materials and Methods
Participant recruiting
Participants were recruited through the InnovATEBIO newsletter (https://innovatebio.org/newsletters) and website (https://InnovATEBIO.org), our website (Antibody-Engineers.org), and announcements by QUBES (QUBES.org) and BioMolViz (BioMolViz.org). We used MailChimp (MailChimp.com) to set up an embedded form in our website so visitors could subscribe and receive emails about registration and updates. We also gave presentations to faculty through InnovATEBIO’s webinar series on ATE projects [26], to students participating in the MNT-CURN research project (DUE 2000281), and to faculty in a weekly teaching discussion hosted by the California Bioscience Workforce Development Hub.
Communication and publishing platforms
We used several Google apps (Google.com) during the events: Google Forms for submitting applications, Google Sheets for reviewing applications and assembling teams, Google Drives for housing collections of materials, Google Slides for presentations, Google Docs for documenting results, and YouTube for sharing videos. Hackathon teams used Slack (Slack.com) for discussions, messaging, and sharing documents. Slack is a communication platform commonly used in the biotech industry and by research labs. We used Zoom (Zoom.com) for meetings with breakout rooms for each team and software demos.
Project records were assembled and published through QUBES (https://qubeshub.org/). QUBES is a content management platform that allows teams to work together and publish their results. Materials from coding-related projects were managed in GitHub (GitHub, https://github.com/AntibodyEngineers).
Machine learning resources and molecular data
We used the NSF-supported Jetstream (https://jetstream-cloud.org/) computing resources for our machine learning project. Jetstream provides eight petaFLOPS of supercomputing power to simplify data analysis, boost discovery, and increase the availability of AI resources. Datasets were from the Oxford Protein Informatics Group (OPIG; opig.stats.ox.ac.uk) CoV-AbDab in addition to the OPIG Ablang ML package. Other molecular data were obtained from the NCBI structure database and IEDB.org. Sequences from SARS-CoV-2 variant spike proteins were obtained from the NCBI. iCn3D and other analysis tools were accessed through the NCBI and IEDB databases.
Laboratory materials
Materials for working with antibodies included a yeast (Saccharomyces cerevisiae) library obtained from Protabit that produced antibodies to the SARS-CoV spike protein, SARS-CoV-2 spike protein (Genscript), protein G magnetic beads (MedChemExpress), a plasmid producing antibodies to GFP (N8his-GFPenhancer-GGGGS4-LaG16) (Addgene), a mouse anti-histidine tag (Bio-Rad), and ELISA reagents (AssayPro and ThermoFisher).
Results from hackathon participants
Hackathon logistics
We hosted two Antibody Engineering hackathons in 2022: the first in January (Thursday, Jan. 13th – Sunday, Jan 16th) and the second in August (Monday, Aug. 8th – Thursday, Aug. 11th). While most of the hackathons were virtual, the Affordable Antibody Engineering project was held in the lab at Los Angeles Pierce College in January and at both LA Pierce College and Pasadena Community College in August. The virtual format made the events cost-effective, with no travel or lodging expenses.
About six weeks before the hackathons began, we reviewed applications, assigned applicants to projects, and set up accounts in Slack. Both hackathons followed similar schedules. Each day began with a full group meeting, followed by guest speakers and software demonstrations. The first day focused on introducing the hackathon, goals, logistics, and team introductions. On the second day, teams presented their project plans, and writers attended a meeting to learn how to document the process and their results. The third day allowed teams to discuss obstacles and seek assistance, with daily meetings for planning, discussion, and coordination. All teams presented their work on the final day.
In January, guest speakers introduced antibodies and discussed course-based undergraduate research experiences, antibody manufacturing, antibody validation, and IEDB. In August, speakers discussed engineering antibodies, engineering antibody-producing cells, and engineering antibody-related careers. Software demonstrations included iCn3D, NextStrain.org, SabDab [20], QUBES, and IEDB.org. We recorded the talks to create a library of materials for future participants (Antibody Engineering Hackathon 2022 Playlist https://www.youtube.com/playlist?list=PLSAXB_etwzD9dl3gN4hgCUtAdspV5FfP9).
A common practice in NIH-sponsored hackathons has been to organize the event around an overarching topic with multiple subprojects [Allissa Dillman, personal communication]. This structure provides a mechanism for rapid prototyping and testing multiple ideas. Including multiple related projects also provides a broader wealth of information sharing since participants hear presentations on all the projects. A last reason for including multiple projects is to make sure the teams don’t get too large. It’s important for everyone on a team to have a specific role that gives them a chance to learn, participate, and contribute without the pressure to compete with others for something to do.
The January hackathon offered five projects, while the August hackathon featured six. Teams consisted of 3 to 7 members. Project selection was influenced by team leader interests, input from our industry advisory board, and suggestions from applicants. During the hackathons, team members choose different roles to facilitate collaboration. Roles like leader, writer, technical support, database expert, researcher, subject matter lead, and quality control are suggested, along with roles like artists, slide makers, and technical writers. Having multiple roles makes it possible for all members to contribute no matter what their experience level. Other hackathons, such as the Bio-IT World 2023 Hackathon, use roles such as: Data scientist/Analyst, Researcher, IT developer, Entrepreneur, Policy Change, and Consultant/Advisor [7].
Participants
Most hackathon participants were community college students and faculty, with others from four-year colleges, universities, high schools, and industry advisory board members (Table 1). For this study, we defined “participants” as people who registered and participated directly on a hackathon team. Over half of the participants were women (59% and 55% in January and August, respectively), and 44% of survey respondents identified as non-white.
Two classes of high school students were also part of the January hackathon. They were unable to participate in August due to a conflict with their schedule. Since they were not registered, were unable to attend all the events and interacted with the group through their teachers, their experience was not directly comparable to other participants, therefore we did not include their survey results in this report.
Table 1. Summary of participant demographics
| Position | Institutional affiliation | Jan 2022 | Aug 2022 |
| Students | Community college | 11 | 17c |
| University | 2 | 1 | |
| High school | (79, 2 classes) a | 2 | |
| Faculty | Community college | 11 | 11c |
| University | 5 | 4 | |
| 4 yr college | 1 | ||
| High school | 4 | 1 | |
| Industry / Research Institute | 5 | 4c | |
| Total registered | 39 | 40 | |
| Gender | Woman | 23 | 22 |
| Man | 15 | 17 | |
| Transgender | 1 | 1 | |
| Race / ethnicity b | Asian | 7 | |
| Black or African American | 4 | ||
| Hispanic or Latino | 5 | ||
| Middle Eastern or North African | 2 | ||
| Multiracial or multiethnic | 1 | ||
| White | 17 | ||
| Answered | 30 | ||
| Skipped | 3 |
What did participants think?
We surveyed participants at the end of each hackathon and obtained 32 responses in January and 30 in August. Participants described positive and negative aspects, identified the top skills they learned, and indicated their interest in continuing to collaborate on their projects. Students were asked if the hackathons allowed them to practice professional skills such as communication, problem-solving, teamwork, and leadership.
Survey respondents from both hackathons listed their favorite aspects in open-ended comments. The most common responses were learning, collaboration, networking, and discovering new resources [Fig. 1]. Working with multi-generational teams with different experience levels was also cited as a positive factor. One participant shared that “My favorite aspect was being able to talk to industry experts and professors with industry experience who could provide real-world experience on each process I didn’t understand.”
About a third of the respondents shared negative aspects (Jan N=12, Aug N=11). Some participants found it challenging to manage their time (Jan N=5/32, Aug N=1/30). Some found QUBES confusing (Jan N=4/32). A few respondents [Jan (2), Aug (3)] felt that they were rushed and trying to complete too much work in too short a time and that the material was more advanced than they expected. Two cited a lack of help, and one was disappointed that there wasn’t a coding component in their project.
What did hackathon participants learn?
We asked participants to identify the top three things they learned during the events. The software (iCn3D, NextStrain, Slack) and databases (IEDB, OPIG) were listed most often with antibodies and related topics next (Fig. 2). Interestingly, teamwork and collaboration skills were also frequently mentioned as areas of learning [Jan (8), Aug (5)].
Respondents from the January (31/32, 97%) and August (32/33, 97%) hackathons said they learned something new. Additionally, 91% (N=29/32) of the respondents from January and 74% from August (23/31) said they learned new things about themselves. Roughly 20% from both hackathons shared comments related to self-efficacy, stating “I can do this” or that “I love working in a lab.” Some participants mentioned discovering potential career paths. Statements to this effect included, “I learned that I could possibly look into protein engineering as a career,” and “I learned that Bioinformatics might be a next career for me after my graduation from college.” At least one student enrolled in a community college biotechnology program after the January hackathon, suggesting a potential role for hackathons in student recruiting.
Many respondents agreed with statements that indicate a sense of belonging in the scientific community, a positive attitude towards collaborative science, and self-efficacy. Over 90% agreed their input was respected, they were more excited about collaborative science and were more confident about participating in future hackathons (Fig. 3). Over 85% worked outside their comfort zones yet stayed with their projects, demonstrating perseverance.
Professional skills
Our primary goal for the hackathons was to develop research projects that can be incorporated into courses. Although hackathons are only a few days, we wondered if the research aspects and collaborative nature of working on the projects might provide similar benefits to students as undergraduate research.
We asked student participants whether they agreed with statements about practicing professional skills during the hackathon (Fig. 4). Students from both hackathons agreed they were able to practice communication, teamwork, and problem-solving. Nearly 70% agreed they had practiced leadership skills. All the students from the January hackathon and 69% from August planned to include the event on their resumés.
Results from hackathon projects
The January (5) and August (6) projects are listed in Table 2. Several projects have been used with classes or for undergraduate research as shown in the last two columns.
Table 2. Hackathon projects and outcomes
| Hackathon | Project | Project Goal | URE | Used in class |
| Jan, Aug | Affordable Antibody Engineering | Develop a low-tech method for screening antibodies. | X | X |
| Jan, Aug | Immune Epitope Database projects | Develop research projects that use IEDB.org. | X | X |
| Jan, Aug | SARS-CoV-2 vs Antibodies | Predict whether antibodies will neutralize SARS-CoV-2 variants. | X | X |
| Jan | Break an Antibody | Develop mutagenesis strategies for disrupting antibody binding. | X | X |
| Jan | iCn3D for Education | Investigate how iCn3D might be used in high school and college. | X | X |
| Aug | Immuno-Zoo | Create a data set of antibody structures for comparing different features. | ||
| Aug | Machine Learning, Artificial Intelligence (AI) and Antibodies | Develop an antibody data set and classroom examples to help students better understand machine learning and AI. | ||
| Aug | Antibody Company Game | Help develop and test a game based on the process of drug development. | X |
Affordable Antibody Engineering
This team explored methods for screening cells to identify high-affinity antibodies. In January, the team used a method where antibodies are displayed on the surface of yeast cells (Saccharomyces cerevisiae). If yeast cells produce high-affinity antibodies, they bind to the antigen and can be captured with iron beads and magnets. Initially, this work focused on identifying antibodies to the SARS-CoV-2 spike protein. In August, the project pivoted to working with nanobodies that bind to Green Fluorescent Protein (GFP). The screening laboratory protocols are currently being developed through undergraduate research by students at LA Pierce College.
Immune Epitope Database (IEDB) projects
This team explored using IEDB to predict good epitopes for creating vaccines and looked at using the Influenza Research Database to find and analyze protein sequences from influenza strains in different parts of the US. They also developed an activity (NetChop, [27]) that uses the IEDB Immunome Browser and an epitope prediction tool, DiscoTope. In the NetChop project, students predict B and T cell epitopes using SARS-CoV-2 as a model. This project is designed to improve student understanding of protein structure and the different responses of B and T cells to epitopes.
Break an Antibody
Our advisory board suggested this project. Successfully engineering an antibody requires modeling the chemical interactions between an antibody and an epitope and evaluating the results of potential changes. Our board suggested it would be easier for students to engineer changes that disrupt binding than it would be to determine if the binding was improved.
In this project, a student identifies key amino acids in the paratope and models the changes in chemical interactions in iCn3D when they are replaced with different amino acids. This process is like the work shown in Fig. 6. The ability of an amino acid substitution to disrupt antibody binding could be tested in vitro by comparing the binding ability of the mutant with the original antibody.
This team used Drugbank.com to identify therapeutic antibodies with available protein sequences and explored three different antibodies: an anti-GFP nanobody and two antibodies that are used as drugs (rituximab and cetuximab). They created drafts of learning objectives and core competencies that would be addressed through these projects.
ImmunoZoo
This team worked to compile a dataset of antibody structures that students could use to compare antibodies from different species. The team located antibodies from llamas and humans but found unanticipated challenges with interpreting database information, making this project more complicated than expected.
SARS-CoV-2 vs. Antibodies
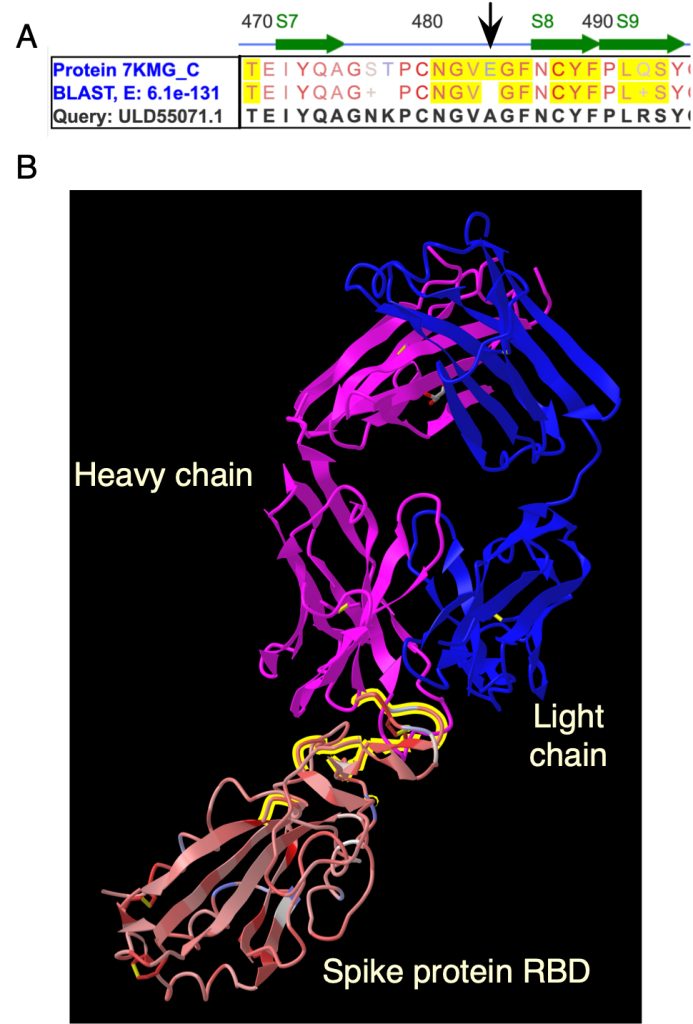
This team developed a CURE based on work from an ISMB hackathon [28], where students would investigate commercially available anti-spike protein antibodies and use iCn3D to determine if they would protect against SARS-CoV-2 variants [29]. Each student has a different antibody. They start by annotating the antibody binding site in iCn3D (Fig. 5B). Then, they find a variant in NextStrain.org. Links from NextStrain to the NCBI are used to get the sequence of the variant spike protein.
The protein BLAST algorithm in iCn3D is used to align the variant sequence to a sequence of an older version of the spike protein bound to an antibody (Fig. 5A). A visual scan of the alignment shows mutations in the antibody binding site. Last, mutation prediction tools in iCn3D are used to model the effects of the mutation on the ability to form chemical interactions with the antibody (Fig. 6A).
In the example (Fig. 5, 6), a student would use the interaction data to create a list of predicted interactions between the original amino acid, in this case, a glutamic acid at position 484 (E484) in the spike protein, and amino acids in the antibody heavy and light chains. They would compare those interactions with the predictions for the A484 variant. Fig. 6A shows that four interactions are potentially lost: a hydrogen bond and contact with R50 (arginine) in the heavy chain, a salt bridge with R96 in the light chain, and a contact with Y101 (tyrosine) in the heavy chain. Losing the two interactions with R50 and the salt bridge, the strongest interaction, is likely to impair binding and allow this variant to escape from this antibody.

iCn3D for Education
This group explored the educational applications of iCn3D for high school and college settings. One activity involved high school seniors using iCn3D to compare two antibody structures and create a Venn diagram noting their similarities and differences. The group also examined how antibody research projects could align with the 5E instruction model [30] (engage, explore, explain, elaborate, evaluate), Next Generation Science Standards [31], and Vision and Change [32]. They compiled datasets of antibodies bound to influenza hemagglutinin and SARS-CoV-2 proteins, as well as structures of Epstein-Barr viral proteins with an antibody.
Machine Learning, Artificial Intelligence, and Antibodies
The project aimed to make machine learning (ML) more accessible and relevant to biotechnology. ML concepts are often communicated at an expert level, using terms like statistical methods, neural nets, convolutional neural nets, and transformers. Moreover, the examples don’t apply to biotechnology, often focusing on sorting dogs, cats, and handwriting samples. Our team chose to train an ML model to analyze the protein sequence of an antibody and determine if it can bind to the receptor binding domain of the SARS-CoV-2 spike protein.
For the hackathon, we set up CPU and GPU virtual machines (VMs) with accounts for 10 team members and installed the Python programming language (Python.org) and relevant Python libraries. The Python libraries included machine learning packages such as TensorFlow for computing and Pandas for data preparation. In addition to the libraries, Jupyter notebook software (Jupyter lab) was installed to give team members a web-based interface to develop and share code.
The team obtained protein sequence datasets from the Oxford Protein Informatics Group (OPIG; opig.stats.ox.ac.uk) – CoV-AbDab, which included 9276 relevant COVID sequences. The OPIG Ablang ML package was used for ML-based sequence analysis.
Some members had computing experience and were able to write scripts to clean data and work with the different Python libraries. However, only one team member had enough programming experience to build and test the ML prediction model. Other members were new to this type of computing and using Jupyter Notebooks, which unexpectedly required a significant amount of instruction time.
Regarding ML prediction, we preprocessed a dataset of 4000 sequences containing variable regions from neutralizing and non-neutralizing anti-SARS-CoV-2 spike protein antibodies into 768 attributes using the AbLang library. A first step in ML is to convert data into numerical n-dimensional vectors so that each datum is unique. The data were split into training data (3200 sequences) and test data (800 sequences). The training data were processed in the artificial neural network to build a model that distinguishes neutralizing antibodies from non-neutralizing. The test data were then used to measure the model’s predictive quality. With 3200 training sequences, our model had a 71% accuracy. Training the model on a 16-core CPU took only five minutes.
Antibody Company Game
The Antibody company game team collaborated on an early version of a game (created through DUE 1764225) and focused on developing online gameplay mechanics. They designed Career and Data cards, established data costs, defined criteria for progressing through drug development phases, and tested the game. The game, Biotechopoly™ Antibody Edition, will serve as an educational tool for introducing biotechnology careers, business concepts, drug development, and reinforcing knowledge of GMPs and GLPs.
Discussion
We learned that hackathons can be an efficient platform for engaging a community in developing prototypes and testing new ideas. At the same time, the short intense nature of these events can be stressful, both for the organizers and the participants. Four of the organizers had participated in at least one hackathon and had some idea of what to expect. Nevertheless, being a hackathon host comes with a different level of responsibility than being a participant. In this section, we will walk through some challenges, discuss changes we made between the two events, and describe changes we will implement in future events.
Our first surprise was in recruiting participants. Over 70 people signed up for the first event, with 18 outside the US. Given our inexperience and small team, we decided to limit acceptances to US applicants. This helped minimize time zone challenges and allowed us to create smaller teams. We also added text to the application form to indicate eligibility.
We were surprised by the high number of student applicants and initially concerned about the dynamics of mixed student-faculty teams. Fortunately, the mixed teams worked remarkably well. The students’ enthusiasm and innovative ideas regarding web technologies pleasantly surprised the faculty. In fact, faculty members considered the collaboration with students to be an unexpected benefit. Students brought great energy to the projects and, in some cases, took the lead in completing most of the work. According to surveys, they also developed a newfound appreciation for faculty efforts in curriculum creation and enjoyed the opportunity to collaborate as peers.
High school students
The most significant challenge arose a week before the first hackathon when we discovered a high school teacher planned to include two classes, totaling 79 seniors. Due to logistical constraints, the high school students had limited participation. School rules prevented the teachers from requiring them to attend during the weekend since those days were outside school hours. The IT policies at the students’ high school also prevented them from using Slack.
To overcome these challenges, the high school teachers agreed to be liaisons to their hackathon teams in Slack and assumed leadership roles for their classes. Although the students couldn’t fully engage in real-time activities, they were able to attend selected talks and daily Zoom sessions, meet with at least one scientist participant, and had access to session recordings. Additionally, Dr. Porter visited one of the high school classes and demonstrated how to find antibodies in the NCBI structure database and analyze them in iCn3D.
An unexpected benefit emerged when Dr. Menshew’s class of 47 high school seniors worked with iCn3D and provided feedback to Dr. Jiyao Wang, iCn3D’s lead developer, and a hackathon participant. The students were thrilled to discover that Dr. Wang incorporated some of their suggestions into the iCn3D program during the event. Furthermore, the high school students completed an assignment comparing different antibodies, which eventually led to the development of the ImmunoZoo project in August.
Communication overload
We learned in January that asking our participants to navigate between Slack, Zoom, QUBES, Google Drive, and Google Docs, in addition to learning GitHub, and the science-focused software and databases (iCn3D, IEDB, SabDab, NextStrain.org, and the NCBI) was too overwhelming. We modified our workflow in August by using a Google Drive specifically for the hackathon with a folder for each project. Instead of directing team members to post in QUBES, we had them use their team folder and provided instructions for using Google Drive and Google apps.
Changes between the first and second hackathons
Using the January survey data and our observations, we made several adjustments to the agenda for the second hackathon. These changes included:
- Asking applicants to agree to the time commitment (8 hours per day).
- Having the event take place during the week and scheduling fewer talks and more breaks.
- Adding team meetings to the schedule to ensure availability and help team leaders.
- Setting up Slack accounts for participants a month in advance with preparatory materials.
Despite these changes, the most common feedback from participants was the need for more background information on antibodies and the projects before the hackathon. Therefore, for our upcoming hackathon, we will provide additional background information and hold an orientation session two weeks prior to the event. This session will increase the likelihood that all participants understand how to use Slack and have the information they need to become familiar with antibodies.
The importance of personnel
We learned that hackathons work best when two crucial roles are filled. The first and most vital role is that of the hackathon manager. This individual possesses knowledge of the hackathon’s timelines and is responsible for setting up the online environment. They answer questions, provide technical support, and guide participants when they face difficulties during the event. Crucially, the hackathon manager meets with the writers from each team and explains the process of documenting their work. This aspect is particularly helpful since having project information summarized and accessible in the designated Google folders makes the projects easier to complete.
The second key role is that of the team lead. Team leads organize team meetings during the hackathon and make sure that everyone has a voice and an opportunity to present. They assist in assigning team roles and managing the project scope, facilitating the team’s ability to achieve results. The significance of team leads was highlighted in August when we attempted to handle too many projects, inadvertently leaving a team of two students without a leader. Although these students persevered, they admitted to feeling lost and frustrated. We learned from this experience and will avoid these situations in the future.
Conclusions
We investigated the use of hackathons as a platform for creating undergraduate research projects, with the accompanying goals of building community and facilitating learning. Through this work, we determined that hackathons are an effective format for achieving these goals. During the two hackathons, we initiated multiple projects. Two projects resulted in curriculum publications in QUBES [27, 29]. Some were paused after a single event, while other projects continued to be developed (Table 2).
In terms of community building, the intense and collaborative nature of the events fostered a shared experience and promoted a sense of community. Participants frequently mentioned learning, collaboration, and networking as highly positive aspects. Almost a quarter of the individuals who participated in the January hackathon returned in August. Moreover, two project teams continued meeting independently over the past year and plan to participate in August 2023.
Survey results indicated that nearly all respondents learned new things, with technical skills being a prominent area of growth. Additionally, participants highlighted learning about teamwork and collaboration. Some expressed surprise at the advanced material but ultimately appreciated the interactions with team members at various levels, demonstrating the value of hackathons as an environment for faculty to prototype new labs and obtain input from a diverse group of team members, from students to scientists.
While our primary focus was creating research projects for undergraduate students, we observed that student participants demonstrated outcomes similar to those attributed to undergraduate research experiences [6]. Their survey responses (Figs 3-4) indicated increased self-efficacy, a sense of belonging, stepping outside of their comfort zones, and enthusiasm for collaborative science, and they could make meaningful contributions. These findings may not be surprising since all the hackathon teams were engaged in short-term research projects. Two notable differences, however, were the short period of time and the focus on collaborative work, as opposed to project ownership. Consequently, hackathons can be a valuable practice for students preparing for careers in the workforce. They offer some of the advantages of undergraduate research while providing a more realistic model of industry practices that prioritize teamwork over individual research.
Acknowledgments. This work was supported by the National Science Foundation (NSF) under award DUE 2055036.
Disclosures. The authors declare no conflicts of interest.
[1] J. S. Krim, L. E. Coté, R. S. Schwartz, E. M. Stone, J. J. Cleeves, K. J. Barry, W. Burgess, S. R. Buxner, J. M. Gerton, L. Horvath, J. M. Keller, S. C. Lee, S. M. Locke, and B. M. Rebar, “Models and Impacts of Science Research Experiences: A Review of the Literature of CUREs, UREs, and TREs,” LSE 18, ar65 (2019).
[2] S. E. Rodenbusch, P. R. Hernandez, S. L. Simmons, and E. L. Dolan, “Early Engagement in Course-Based Research Increases Graduation Rates and Completion of Science, Engineering, and Mathematics Degrees,” LSE 15, ar20 (2016).
[3] M. Estrada, M. Burnett, A. G. Campbell, P. B. Campbell, W. F. Denetclaw, C. G. Gutiérrez, S. Hurtado, G. H. John, J. Matsui, R. McGee, C. M. Okpodu, T. J. Robinson, M. F. Summers, M. Werner-Washburne, and M. Zavala, “Improving Underrepresented Minority Student Persistence in STEM,” LSE 15, es5 (2016).
[4] AACC “Fast Facts 2023,” https://www.aacc.nche.edu/research-trends/fast-facts/.
[5] “Course-based Undergraduate Research Experiences: Current knowledge and future directions,” https://sites.nationalacademies.org/cs/groups/dbassesite/documents/webpage/dbasse_177288.pdf.”
[6] C. T. Leonetti, H. Lindberg, D. O. Schwake, and R. L. Cotter, “A Call to Assess the Impacts of Course-Based Undergraduate Research Experiences for Career and Technical Education, Allied Health, and Underrepresented Students at Community Colleges,” LSE 22, ar4 (2023).
[7] “Hackathon – Bio-IT World Conference & Expo,” https://www.bio-itworldexpo.com/fair-data-hackathon.
[8] “Codeathon – Tools for Shareable Protein Analysis,” https://www.iscb.org/ismb2022-program/protein-codeathon.
[9] NNLM “Love Data Week 2023 – NCDS Bit by Bit,” (2023).
[10] G. C. Ferreira, J. Oberstaller, R. Fonseca, T. E. Keller, S. R. Adapa, J. Gibbons, C. Wang, X. Liu, C. Li, M. Pham, G. W. Dayhoff Ii, L. M. Duong, L. T. Reyes, L. E. Laratelli, D. Franz, S. Fatumo, A. G. Bari, A. Freischel, L. Fiedler, O. Dokur, K. Sharma, D. Cragun, B. Busby, and R. H. Y. Jiang, “Iron Hack – A symposium/hackathon focused on porphyrias, Friedreich’s ataxia, and other rare iron-related diseases,” F1000Res 8, 1135 (2019).
[11] C. for D. and R. Health, “Guidance for Over-the-Counter (OTC) Human Chorionic Gonadotropin (hCG) 510(k)s – Guidance for Industry and FDA Reviewers/Staff,” https://www.fda.gov/regulatory-information/search-fda-guidance-documents/guidance-over-counter-otc-human-chorionic-gonadotropin-hcg-510ks-guidance-industry-and-fda.
[12] C. for D. and R. Health, “At-Home OTC COVID-19 Diagnostic Tests,” FDA (2023).
[13] “Antibody therapeutics approved or in regulatory review in the EU or US,” https://www.antibodysociety.org/resources/approved-antibodies/.
[14] Z. Wang, G. Wang, H. Lu, H. Li, M. Tang, and A. Tong, “Development of therapeutic antibodies for the treatment of diseases,” Mol Biomed 3, 35 (2022).
[15] P. T. Jones, P. H. Dear, J. Foote, M. S. Neuberger, and G. Winter, “Replacing the complementarity-determining regions in a human antibody with those from a mouse,” Nature 321, 522–525 (1986).
[16] N. Tsurushita, P. R. Hinton, and S. Kumar, “Design of humanized antibodies: From anti-Tac to Zenapax,” Methods 36, 69–83 (2005).
[17] F. V. Suurs, M. N. Lub-de Hooge, E. G. E. De Vries, and D. J. A. De Groot, “A review of bispecific antibodies and antibody constructs in oncology and clinical challenges,” Pharmacology & Therapeutics 201, 103–119 (2019).
[18] J. Wang, P. Youkharibache, D. Zhang, C. J. Lanczycki, R. C. Geer, T. Madej, L. Phan, M. Ward, S. Lu, G. H. Marchler, Y. Wang, S. H. Bryant, L. Y. Geer, and A. Marchler-Bauer, “iCn3D, a web-based 3D viewer for sharing 1D/2D/3D representations of biomolecular structures,” Bioinformatics 36, 131–135 (2020).
[19] M. Mendes, J. Mahita, N. Blazeska, J. Greenbaum, B. Ha, K. Wheeler, J. Wang, D. Shackelford, A. Sette, and B. Peters, “IEDB‐3D 2.0: Structural data analysis within the Immune Epitope Database,” Protein Science 32, e4605 (2023).
[20] J. Dunbar, K. Krawczyk, J. Leem, C. Marks, J. Nowak, C. Regep, G. Georges, S. Kelm, B. Popovic, and C. M. Deane, “SAbPred: a structure-based antibody prediction server,” Nucleic Acids Res 44, W474–W478 (2016).
[21] “Addgene: Antibody Plasmid Collection,” https://www.addgene.org/collections/antibody-plasmids/.
[22] “NISTCHO,” NIST (2016).
[23] S. Muyldermans, “Nanobodies: Natural Single-Domain Antibodies,” Annu. Rev. Biochem. 82, 775–797 (2013).
[24] W. R. Strohl and L. M. Strohl, eds., “18 – Cell line development,” in Therapeutic Antibody Engineering, Woodhead Publishing Series in Biomedicine (Woodhead Publishing, 2012), pp. 421–595.
[25] “The InnovATEBIO Network,” https://innovatebio.org/.
[26] “Spring 2023 InnoATEBIO ATE Project Talks Webinar Series,” https://innovatebio.org/event/spring-2023-innovatebio-ate-project-talks-webinar-series.
[27] M. Mallari, S. Vemu, V. Carson, G. Afshar, A. Estrada, C. Wilson, and S. Porter, “Using the IEDB to Predict Proteasomal Cleavage Events and T-cell Epitopes,” (2022).
[28] S. G. Porter and T. M. Smith, “Combining iCn3D and NextStrain to create a novel undergraduate research experience around SARS-CoV-2 variants and commercial antibodies,” Frontiers in Genetics 14, (2023).
[29] S. Porter, U. Hilgert, E. Lannan, and M. Stieber, “Evaluating the potential for immune escape: how likely is an antibody to protect against a specific SARS-CoV-2 variant?” (2022).
[30] R. W. Bybee, The BSCS 5E Instructional Model: Creating Teachable Moments (NSTA Press, National Science Teachers Association, 2015).
[31] “Home Page | Next Generation Science Standards,” https://www.nextgenscience.org/.
[32] S. E. Brownell, S. Freeman, M. P. Wenderoth, and A. J. Crowe, “BioCore Guide: A Tool for Interpreting the Core Concepts of Vision and Change for Biology Majors,” LSE 13, 200–211 (2014).